Customize results table
Click on the to add to 'Columns to be displayed' and to the 'Miscellaneous' section below for future use.
Drag and drop to re-order.
Your basket is currently empty.
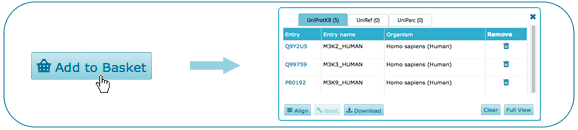
Select item(s) and click on "Add to basket" to create your own collection here
(400 entries max)
Help Results
| Help search results |
|---|
| Accession Provides a stable way of identifying UniProtKB entries |
| Active site Amino acid(s) directly involved in the activity of an enzyme |
| Allergenic properties Information relevant to allergenic proteins |
| Alternative products Description of the different proteins generated from the same gene |
| Annotation score Annotation scores provide a measure of how much annotation has been associated with a given entry or proteome. |
| Binary interactions Information relevant to binary protein-protein interactions |
| Binding site Binding site for any chemical group (co-enzyme, prosthetic group, etc.) |
| Biophysicochemical properties Description of biophysical and physicochemical properties |
| Biotechnological use The use of a specific protein in the biotechnological industry |
| Calcium binding Denotes the position(s) of calcium binding region(s) within the protein |
| Glycosylation Covalently attached glycan group(s) |
| Catalytic activity Description of the reaction(s) catalyzed by an enzyme |
| Caution Warning about possible errors and/or grounds for confusion |
| Chain Extent of a polypeptide chain in the mature protein |
| Cofactor Description of non-protein substance required by an enzyme to be active |
| Coiled coil Denotes the positions of regions of coiled coil within the protein |
| Compositional bias Region of compositional bias in the protein |
| Sequence conflict Description of sequence discrepancies of unknown origin |
| Cross-references section Cross-references that point to data collections other than UniProtKB |
| Cross-link Residues participating in covalent linkage(s) between proteins |
| Developmental stage Description of how the expression of a gene varies according to the stage of cell, tissue or organism development |
| Disruption phenotype Description of the effects caused by the disruption of a protein-encoding gene. |
| Disulfide bond Cysteine residues participating in disulfide bonds |
| DNA binding Denotes the position and type of a DNA-binding domain |
| Domain Denotes the position and type of each modular protein domain |
| Domain Description of the domain(s) present in a protein |
| Encoded on Organelle or plasmid gene source |
| Entry history Dates of creation and modification and version information |
| Entry information section Content of the 'Entry information' section |
| Entry name Mnemonic identifier for a UniProtKB entry |
| Entry status Indicates if the entry has been manually reviewed (UniProtKB/Swiss-Prot) or not (UniProtKB/TrEMBL) |
| Enzyme regulation Description of an enzyme regulatory mechanism |
| Evidences
|
| Expression section Content of the 'Expression' section |
| Family and Domains section Content of the 'Family and Domains' section |
| Function General description of the function(s) of a protein |
| Function section Content of the 'Function' section |
| Gene names Name(s) of the gene(s) that code for the protein |
| Gene Ontology (GO) Selection of Gene Ontology (GO) terms |
| General annotation (Comments) Position-independent annotation |
| Helix Helical regions within the experimentally determined protein structure |
| Induction Description of the effects of environmental factors on the gene expression |
| Initiator methionine Cleaved initiator methionine |
| Interaction section Content of the 'Interaction' section |
| Intramembrane Extent of a region located in a membrane without crossing it |
| Involvement in disease Description of the disease(s) associated with the deficiency of a protein |
| Lipidation Covalently attached lipid group(s) |
| Mass spectrometry Information derived from mass spectrometry experiments |
| Metal binding Binding site for a metal ion |
| Miscellaneous Any relevant information that doesn't fit in any other defined sections |
| Miscellaneous section Content of the 'Miscellaneous' section |
| Modified residue Modified residues excluding lipids, glycans and protein crosslinks |
| Motif Short (up to 20 amino acids) sequence motif of biological interest |
| Mutagenesis Site which has been experimentally altered by mutagenesis |
| Names and Taxonomy section Content of the 'Names and Taxonomy' section |
| Non-adjacent residues Indicates that two residues in a sequence are not consecutive |
| Non-experimental qualifiers Non-experimental qualifiers indicate that the information given is not based on experimental findings. |
| Non-standard residue Occurence of non-standard amino acids (selenocysteine and pyrrolysine) in the protein sequence |
| Non-terminal residue The sequence is incomplete. Indicate that a residue is not the terminal residue of the complete protein. |
| Nucleotide binding Nucleotide phosphate binding region |
| Organism Name of the source organism |
| Pathology and Biotech section Content of the 'Pathology and Biotech' section |
| Pathway Description of associated metabolic pathways |
| Peptide Extent of an active peptide in the mature protein |
| Pharmaceutical use Description of the use of a protein as a pharmaceutical drug |
| Polymorphism Description of polymorphism(s) |
| Post-translational modification Description of post-translational modifications |
| Propeptide Part of a protein that is cleaved during maturation or activation |
| Protein existence Level of evidence that supports the existence of the protein concerned |
| Protein names Name and synonyms of the protein |
| Proteome component Genomic component encoding a set of proteins in a proteome. |
| Proteome identifier Unique identifier assigned to the set of proteins that constitute the proteome. |
| Proteomes Sets of proteins thought to be expressed by organisms whose genomes have been completely sequenced. |
| PTM / Processing section Content of the 'PTM / Processing' section |
| Publications section Literature citations |
| Region Region of interest in the sequence |
| Repeat Denotes the positions of repeated sequence motifs or repeated domains |
| RNA editing Description of amino acid change(s) due to RNA editing |
| Sequence annotation (Features) Position-dependent annotation |
| Sequence caution Warning about possible errors related to the protein sequence |
| Sequence length Sequence length |
| Sequence processing Indicates if the mature form of a protein is derived by processing of a precursor |
| Sequence similarities Description of the sequence similaritie(s) with other proteins |
| Sequence status Indicates if the protein sequence is complete or not |
| Sequences Protein sequence data for all described protein isoforms |
| Sequence section Content of the 'Sequence' section |
| Signal peptide Sequence targeting proteins to the secretory pathway or periplasmic space |
| Similar Proteins section Content of the 'Similar Proteins' section |
| Site Any interesting single amino acid site on the sequence |
| Beta strand Beta strand regions within the experimentally determined protein structure |
| Structure section Content of the 'Structure' section |
| Subcellular location Description of the subcellular location of the mature protein |
| Subcellular location section Content of the 'Subcellular location' section |
| Subunit structure Description of the quaternary structure of a protein |
| Taxonomic identifier NCBI unique identifier for the source organism |
| Taxonomic lineage Taxonomic classification of the source organism |
| Tissue specificity Description of the expression of a gene in different tissues |
| Topological domain Location of non-membrane regions of membrane-spanning proteins |
| Toxic dose Lethal, paralytic, or effect dose, or lethal concentration of a toxin |
| Transit peptide Extent of a transit peptide for organelle targeting |
| Transmembrane Extent of a membrane-spanning region |
| Turn Turns within the experimentally determined protein structure |
| UniProtKB manual Table of contents of the UniProtKB manual |
| Sequence uncertainty Regions of uncertainty in the sequence |
| Alternative sequence Amino acid change(s) producing alternate protein isoforms |
| Natural variant Description of a natural variant of the protein |
| Virus host The species that can be infected by a specific virus |
| Web resources Links to related web resource(s) or database(s) |
| Zinc finger Denotes the position(s) and type(s) of zinc fingers within the protein |

