Summary: Protein kinase domain
Pfam includes annotations and additional family information from a range of different sources. These sources can be accessed via the tabs below.
This is the Wikipedia entry entitled "Protein kinase domain". More...
Protein kinase domain Edit Wikipedia article
| Protein kinase domain | |||
|---|---|---|---|
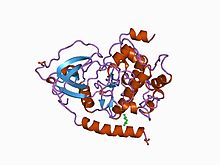
Structure of the catalytic subunit of cAMP-dependent protein kinase.[1]
|
|||
| Identifiers | |||
| Symbol | Pkinase | ||
| Pfam | PF00069 | ||
| InterPro | IPR000719 | ||
| SMART | TyrKc | ||
| PROSITE | PDOC00629 | ||
| SCOP | 1apm | ||
| SUPERFAMILY | 1apm | ||
| OPM superfamily | 417 | ||
| OPM protein | 2w5a | ||
| CDD | cd00180 | ||
|
|||
The protein kinase domain is a structurally conserved protein domain containing the catalytic function of protein kinases.[2][3][4] Protein kinases are a group of enzymes that move a phosphate group onto proteins, in a process called phosphorylation. This functions as an on/off switch for many cellular processes, including metabolism, transcription, cell cycle progression, cytoskeletal rearrangement and cell movement, apoptosis, and differentiation. They also function in embryonic development, physiological responses, and in the nervous and immune system. Abnormal phosphorylation causes many human diseases, including cancer, and drugs that affect phosphorylation can treat those diseases.[5]
Protein kinases possess a catalytic subunit which transfers the gamma phosphate from nucleoside triphosphates (often ATP) to one or more amino acid residues in a protein substrate side chain, resulting in a conformational change affecting protein function. These enzymes fall into two broad classes, characterised with respect to substrate specificity: serine/threonine specific and tyrosine specific.[6]
Contents
[hide]Function
Protein kinase function has been evolutionarily conserved from Escherichia coli to Homo sapiens. Protein kinases play a role in a multitude of cellular processes, including division, proliferation, apoptosis, and differentiation.[7] Phosphorylation usually results in a functional change of the target protein by changing enzyme activity, cellular location, or association with other proteins.
Structure
The catalytic subunits of protein kinases are highly conserved, and several structures have been solved,[8] leading to large screens to develop kinase-specific inhibitors for the treatments of a number of diseases.[9]
Eukaryotic protein kinases[2][3][10][11] are enzymes that belong to a very extensive family of proteins which share a conserved catalytic core common with both serine/threonine and tyrosine protein kinases. There are a number of conserved regions in the catalytic domain of protein kinases. In the N-terminal extremity of the catalytic domain there is a glycine-rich stretch of residues in the vicinity of a lysine residue, which has been shown to be involved in ATP binding. In the central part of the catalytic domain there is a conserved aspartic acid residue which is important for the catalytic activity of the enzyme.[12]
Examples
The following is a list of human proteins containing the protein kinase domain:[13]
AAK1; ABL1; ABL2; ACVR1; ACVR1B; ACVR1C; ACVR2A; ACVR2B; ACVRL1; ADCK1; ADCK2; ADCK3; ADCK4; ADCK5; ADRBK1; ADRBK2; AKT1; AKT2; AKT3; ALPK1; ALPK2; ALPK3; STRADB; CDK15; AMHR2; ANKK1; ARAF; ATM; ATR; AURKA; AURKB; AURKC; AXL; BCKDK; BLK; BMP2K; BMPR1A; BMPR1B; BMPR2; BMX; BRAF; BRSK1; BRSK2; BTK; BUB1; C21orf7; CALM1; CALM2; CALM3; CAMK1; CAMK1D; CAMK1G; CAMK2A; CAMK2B; CAMK2D; CAMK2G; CAMK4; CAMKK1; CAMKK2; CAMKV; CASK; CDK20; CDK1; CDK11B; CDK11A; CDK13; CDK19; CDC42BPA; CDC42BPB; CDC42BPG; CDC7; CDK10; CDK2; CDK3; CDK4; CDK5; CDK6; CDK7; CDK8; CDK9; CDK12; CDK14; CDK16; CDK17; CDK18; CDKL1; CDKL2; CDKL3; CDKL4; CDKL5; CHEK1; CHEK2; CHUK; CIT; CKB; CKM; CLK1; CLK2; CLK3; CLK4; CSF1R; CSK; CSNK1A1; CSNK1A1L; CSNK1D; CSNK1E; CSNK1G1; CSNK1G2; CSNK1G3; CSNK2A1; CSNK2A2; DAPK1; DAPK2; DAPK3; DCLK1; DCLK2; DCLK3; DDR1; DDR2; DMPK; DYRK1A; DYRK1B; DYRK2; DYRK3; DYRK4; EGFR; EIF2AK1; EIF2AK2; EIF2AK3; EIF2AK4; ELK1; EPHA1; EPHA2; EPHA3; EPHA4; EPHA5; EPHA6; EPHA7; EPHA8; EPHB1; EPHB2; EPHB3; EPHB4; ERBB2; ERBB3; ERBB4; ERN1; ERN2; FER; FES; FGFR1; FGFR2; FGFR3; FGFR4; FGR; FLT1; FLT3; FLT4; FYN; GAK; GRK1; GRK4; GRK5; GRK6; GRK7; GSK3A; GSK3B; GUCY2C; GUCY2D; GUCY2E; GUCY2F; HCK; HIPK1; HIPK2; HIPK3; HIPK4; HUNK; ICK; IGF1R; IGF2R; IKBKB; IKBKE; ILK; INSR; IRAK1; IRAK2; IRAK3; IRAK4; ITK; JAK1; JAK2; JAK3; KALRN; KDR; SIK3; KSR2; LATS1; LATS2; LIMK1; LCK; LIMK2; LRRK1; LRRK2; LYN; MAK; MAP2K1; MAP2K2; MAP2K3; MAP2K4; MAP2K5; MAP2K6; MAP2K7; MAP3K1; MAP3K10; MAP3K11; MAP3K12; MAP3K13; MAP3K14; MAP3K15; MAP3K2; MAP3K3; MAP3K4; MAP3K5; MAP3K6; MAP3K7; MAP3K8; MAP3K9; MAP4K1; MAP4K2; MAP4K3; MAP4K4; MAP4K5; MAPK1; MAPK10; MAPK12; MAPK13; MAPK14; MAPK15; MAPK3; MAPK4; MAPK6; MAPK7; MAPK8; MAPK9; MAPKAPK2; MAPKAPK3; MAPKAPK5; MARK1; MARK2; MARK3; MARK4; MAST1; MAST2; MAST3; MAST4; MASTL; MELK; MERTK; MET; MINK1; MKNK1; MKNK2; MLKL; MOS; MST1R; MST4; MTOR; MYLK; MYLK2; MYLK3; MYLK4; NEK1; NEK10; NEK11; NEK2; NEK3; NEK4; NEK5; LOC100506859; NEK6; NEK7; NEK8; NEK9; MGC42105; NLK; NRK; NTRK1; NTRK2; NTRK3; NUAK1; NUAK2; OBSCN; OXSR1; PAK1; PAK2; PAK3; PAK4; PAK6; PAK7; PASK; PBK; PDGFRA; PDGFRB; PDIK1L; PDPK1; PHKA1; PHKB; PHKG1; PHKG2; PIK3R4; PIM1; PIM2; PIM3; PINK1; PKMYT1; PKN1; PKN2; PKN3; PLK1; PLK2; PLK3; PLK4; PNCK; PRKAA1; PRKAA2; PRKACA; PRKACB; PRKACG; PRKCA; PRKCB; PRKCD; PRKCE; PRKCG; PRKCH; PRKCI; PRKCQ; PRKCZ; PRKD1; PRKD2; PRKD3; PRKG1; PRKG2; PRKX; LOC389906; PRKY; PRPF4B; PSKH1; PSKH2; PTK2; PTK2B; RAF1; RAGE; RET; RIP3; RIPK1; RIPK2; RIPK3; RIPK4; ROCK1; ROCK2; ROR1; ROR2; ROS1; RPS6KA1; RPS6KA2; RPS6KA3; RPS6KA4; RPS6KA5; RPS6KA6; RPS6KB1; RPS6KB2; RPS6KC1; RPS6KL1; RYK; SCYL1; SCYL2; SCYL3; SGK1; LOC100130827; SGK196; SGK2; SGK3; SGK494; SIK1; SIK2; SLK; SNRK; SPEG; SRC; SRPK1; SRPK2; SRPK3; STK10; STK11; STK16; STK17A; STK17B; STK19; STK24; STK25; STK3; STK31; STK32A; STK32B; STK32C; STK33; STK35; STK36; STK38; STK38L; STK39; STK4; STK40; SYK; TAOK1; TAOK2; TAOK3; TBCK; TBK1[disambiguation needed]; TEC; TESK1; TESK2; TGFBR1; TGFBR2; TIE1; TIE2; TLK1; TLK2; TNIK; TNK1; TNK2; TSSK1B; TSSK2; TSSK3; TSSK4; TTBK1; TTBK2; TTK; TWF2; TXK; TYK2; TYRO3; UHMK1; ULK1; ULK2; ULK3; ULK4; VRK1; VRK2; VRK3; WEE1; WEE2; WNK1; WNK2; WNK3; WNK4; YES1; ZAK; ZAP70;
References
- ^ Knighton DR, Bell SM, Zheng J, et al. (May 1993). "2.0 A refined crystal structure of the catalytic subunit of cAMP-dependent protein kinase complexed with a peptide inhibitor and detergent". Acta Crystallogr. D Biol. Crystallogr. 49 (Pt 3): 357–61. doi:10.1107/S0907444993000502. PMID 15299526.
- ^ a b Hanks SK, Quinn AM (1991). [2] Protein kinase catalytic domain sequence database: Identification of conserved features of primary structure and classification of family members. "Protein kinase catalytic domain sequence database: identification of conserved features of primary structure and classification of family members". Meth. Enzymol. Methods in Enzymology 200: 38–62. doi:10.1016/0076-6879(91)00126-H. ISBN 978-0-12-182101-2. PMID 1956325.
- ^ a b Hanks SK, Hunter T (May 1995). "Protein kinases 6. The eukaryotic protein kinase superfamily: kinase (catalytic) domain structure and classification". FASEB J. 9 (8): 576–96. PMID 7768349.
- ^ Scheeff ED, Bourne PE (October 2005). "Structural evolution of the protein kinase-like superfamily". PLoS Comput. Biol. 1 (5): e49. doi:10.1371/journal.pcbi.0010049. PMC 1261164. PMID 16244704.
- ^ G. Manning, D. B. Whyte, R. Martinez, T. Hunter, S. Sudarsanam, The Protein Kinase Complement of the Human Genome, Science 6 December 2002, 298:1912-1934 doi:10.1126/science.1075762
- ^ Hunter T, Hanks SK, Quinn AM (1988). "The protein kinase family: conserved features and deduced phylogeny of the catalytic domains". Science 241 (4861): 42–51. doi:10.1126/science.3291115. PMID 3291115.
- ^ Manning G, Plowman GD, Hunter T, Sudarsanam S (October 2002). "Evolution of protein kinase signaling from yeast to man". Trends Biochem. Sci. 27 (10): 514–20. doi:10.1016/S0968-0004(02)02179-5. PMID 12368087.
- ^ Stout TJ, Foster PG, Matthews DJ (2004). "High-throughput structural biology in drug discovery: protein kinases". Curr. Pharm. Des. 10 (10): 1069–82. doi:10.2174/1381612043452695. PMID 15078142.
- ^ Li B, Liu Y, Uno T, Gray N (August 2004). "Creating chemical diversity to target protein kinases". Comb. Chem. High Throughput Screen. 7 (5): 453–72. doi:10.2174/1386207043328580. PMID 15320712.
- ^ Hanks SK (2003). "Genomic analysis of the eukaryotic protein kinase superfamily: a perspective". Genome Biol. 4 (5): 111. doi:10.1186/gb-2003-4-5-111. PMC 156577. PMID 12734000.[dead link]
- ^ Hunter T (1991). [1] Protein kinase classification. "Protein kinase classification". Meth. Enzymol. Methods in Enzymology 200: 3–37. doi:10.1016/0076-6879(91)00125-G. ISBN 978-0-12-182101-2. PMID 1835513.
- ^ Knighton DR, Zheng JH, Ten Eyck LF, Ashford VA, Xuong NH, Taylor SS, Sowadski JM (July 1991). "Crystal structure of the catalytic subunit of cyclic adenosine monophosphate-dependent protein kinase". Science 253 (5018): 407–14. doi:10.1126/science.1862342. PMID 1862342.
- ^ Manning G, Whyte DB, Martinez R, Hunter T, Sudarsanam S (December 2002). "The protein kinase complement of the human genome". Science 298 (5600): 1912–34. doi:10.1126/science.1075762. PMID 12471243.
External links
- Eukaryotic Linear Motif resource motif class LIG_AGCK_PIF_1
- Eukaryotic Linear Motif resource motif class LIG_AGCK_PIF_2
- Eukaryotic Linear Motif resource motif class LIG_AGCK_PIF_3
- Eukaryotic Linear Motif resource motif class LIG_MAPK_1
- Eukaryotic Linear Motif resource motif class LIG_MAPK_2
- Eukaryotic Linear Motif resource motif class MOD_CDK_1
- Eukaryotic Linear Motif resource motif class MOD_CK1_1
- Eukaryotic Linear Motif resource motif class MOD_CK2_1
- Eukaryotic Linear Motif resource motif class MOD_GSK3_1
- Eukaryotic Linear Motif resource motif class MOD_LATS_1
- Eukaryotic Linear Motif resource motif class MOD_PK_1
- Eukaryotic Linear Motif resource motif class MOD_PKA_1
- Eukaryotic Linear Motif resource motif class MOD_PKA_2
- Eukaryotic Linear Motif resource motif class MOD_PKB_1
- Eukaryotic Linear Motif resource motif class MOD_ProDKin_1
- Eukaryotic Linear Motif resource motif class MOD_TYR_DYR
This page is based on a Wikipedia article. The text is available under the Creative Commons Attribution/Share-Alike License.
This tab holds the annotation information that is stored in the Pfam database. As we move to using Wikipedia as our main source of annotation, the contents of this tab will be gradually replaced by the Wikipedia tab.
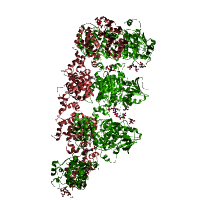
Example structure
PDB entry 3SDM: Structure of oligomeric kinase/RNase Ire1 in complex with an oligonucleotideProtein kinase domain Provide feedback
No Pfam abstract.
Literature references
-
Hanks SK, Quinn AM; , Methods Enzymol 1991;200:38-62.: Protein kinase catalytic domain sequence database: identification of conserved features of primary structure and classification of family members. PUBMED:1956325 EPMC:1956325
-
Hanks SK, Hunter T; , FASEB J 1995;9:576-596.: Protein kinases 6. The eukaryotic protein kinase superfamily: kinase (catalytic) domain structure and classification. PUBMED:7768349 EPMC:7768349
-
Hunter T, Plowman GD; , Trends Biochem Sci 1997;22:18-22.: The protein kinases of budding yeast: six score and more. PUBMED:9020587 EPMC:9020587
Internal database links
| Similarity to PfamA using HHSearch: | RIO1 APH Kdo Pkinase_Tyr YrbL-PhoP_reg Kinase-like |
External database links
| HOMSTRAD: | kinase TyrKc |
| PANDIT: | PF00069 |
| PRINTS: | PR00109 |
| PROSITE: | PDOC00100 PDOC00212 PDOC00213 PDOC00629 |
| PROSITE profile: | PS50011 |
| Pseudofam: | PF00069 |
| SCOP: | 1apm |
| SMART: | STYKc S_TKc TyrKc |
| SYSTERS: | Pkinase |
This tab holds annotation information from the InterPro database.
InterPro entry IPR000719
Protein phosphorylation, which plays a key role in most cellular activities, is a reversible process mediated by protein kinases and phosphoprotein phosphatases. Protein kinases catalyse the transfer of the gamma phosphate from nucleotide triphosphates (often ATP) to one or more amino acid residues in a protein substrate side chain, resulting in a conformational change affecting protein function. Phosphoprotein phosphatases catalyse the reverse process. Protein kinases fall into three broad classes, characterised with respect to substrate specificity [PUBMED:3291115]:
- Serine/threonine-protein kinases
- Tyrosine-protein kinases
- Dual specificity protein kinases (e.g. MEK - phosphorylates both Thr and Tyr on target proteins)
Protein kinase function is evolutionarily conserved from Escherichia coli to human [PUBMED:12471243]. Protein kinases play a role in a multitude of cellular processes, including division, proliferation, apoptosis, and differentiation [PUBMED:12368087]. Phosphorylation usually results in a functional change of the target protein by changing enzyme activity, cellular location, or association with other proteins. The catalytic subunits of protein kinases are highly conserved, and several structures have been solved [PUBMED:15078142], leading to large screens to develop kinase-specific inhibitors for the treatments of a number of diseases [PUBMED:15320712].
Eukaryotic protein kinases [PUBMED:12734000, PUBMED:7768349, PUBMED:1835513, PUBMED:1956325, PUBMED:3291115] are enzymes that belong to a very extensive family of proteins which share a conserved catalytic core common with both serine/threonine and tyrosine protein kinases. There are a number of conserved regions in the catalytic domain of protein kinases. In the N-terminal extremity of the catalytic domain there is a glycine-rich stretch of residues in the vicinity of a lysine residue, which has been shown to be involved in ATP binding. In the central part of the catalytic domain there is a conserved aspartic acid residue which is important for the catalytic activity of the enzyme [PUBMED:1862342].
This entry represents the protein kinase catalytic domain found in serine/threonine-protein kinases, tyrosine-protein kinases and dual specificity protein kinases.
Gene Ontology
The mapping between Pfam and Gene Ontology is provided by InterPro. If you use this data please cite InterPro.
| Molecular function | ATP binding (GO:0005524) |
| protein kinase activity (GO:0004672) | |
| Biological process | protein phosphorylation (GO:0006468) |
Domain organisation
Below is a listing of the unique domain organisations or architectures in which this domain is found. More...
There are 70220 sequences with the following architecture: Pkinase
Q9YH61_DANRE [Danio rerio (Zebrafish) (Brachydanio rerio)] Uncharacterized protein (440 residues) Show all sequences with this architecture.There are 3505 sequences with the following architecture: Pkinase x 2
SRPK3_MOUSE [Mus musculus (Mouse)] SRSF protein kinase 3 EC=2.7.11.1 (565 residues) Show all sequences with this architecture.There are 1200 sequences with the following architecture: Pkinase, Pkinase_C
Q9W6Y9_XENLA [Xenopus laevis (African clawed frog)] Kinase (501 residues) Show all sequences with this architecture.There are 1157 sequences with the following architecture: Pkinase, PASTA x 3
Q0PIH7_HELMO [Heliobacillus mobilis] Ser/Thr protein kinase (634 residues) Show all sequences with this architecture.There are 692 sequences with the following architecture: Lectin_legB, Pkinase
Q0DY65_ORYSJ [Oryza sativa subsp. japonica (Rice)] Os02g0712700 protein (747 residues) Show all sequences with this architecture.There are 689 sequences with the following architecture: Pkinase, PASTA x 4
Q1B026_RUBXD [Rubrobacter xylanophilus (strain DSM 9941 / NBRC 16129)] Serine/threonine protein kinase (666 residues) Show all sequences with this architecture.There are 683 sequences with the following architecture: PBD, Pkinase
STE20_YEAST [Saccharomyces cerevisiae (strain ATCC 204508 / S288c) (Baker's yeast)] Serine/threonine-protein kinase STE20 EC=2.7.11.1 (939 residues) Show all sequences with this architecture.There are 680 sequences with the following architecture: Pkinase, CNH
M4K4_MOUSE [Mus musculus (Mouse)] Mitogen-activated protein kinase kinase kinase kinase 4 EC=2.7.11.1 (1233 residues) Show all sequences with this architecture.There are 639 sequences with the following architecture: Pkinase, KA1
Q9ZRJ3_ORYSA [Oryza sativa (Rice)] OSK4 (509 residues) Show all sequences with this architecture.There are 625 sequences with the following architecture: Pkinase, NAF
CIPKG_ARATH [Arabidopsis thaliana (Mouse-ear cress)] CBL-interacting serine/threonine-protein kinase 16 EC=2.7.11.1 (469 residues) Show all sequences with this architecture.There are 514 sequences with the following architecture: Activin_recp, TGF_beta_GS, Pkinase
Q90754_CHICK [Gallus gallus (Chicken)] Protein kinase (533 residues) Show all sequences with this architecture.There are 483 sequences with the following architecture: FHA, Pkinase
Q761W8_ENTHI [Entamoeba histolytica] Protein kinase domain containing protein (398 residues) Show all sequences with this architecture.There are 453 sequences with the following architecture: Pkinase, CaMKII_AD
Q8IWE0_HUMAN [Homo sapiens (Human)] Calcium/calmodulin-dependent protein kinase II alpha (478 residues) Show all sequences with this architecture.There are 431 sequences with the following architecture: Pkinase, Pkinase_C, Pkinase
KS6A5_HUMAN [Homo sapiens (Human)] Ribosomal protein S6 kinase alpha-5 EC=2.7.11.1 (802 residues) Show all sequences with this architecture.There are 415 sequences with the following architecture: Pkinase, OSR1_C
WNK4_HUMAN [Homo sapiens (Human)] Serine/threonine-protein kinase WNK4 EC=2.7.11.1 (1243 residues) Show all sequences with this architecture.There are 369 sequences with the following architecture: RGS, Pkinase
Q9U756_HOMAM [Homarus americanus (American lobster)] G protein-coupled receptor kinase type 2 (690 residues) Show all sequences with this architecture.There are 364 sequences with the following architecture: B_lectin, S_locus_glycop, PAN_2, Pkinase
Q851M7_ORYSJ [Oryza sativa subsp. japonica (Rice)] cDNA clone:002-179-G09, full insert sequence (858 residues) Show all sequences with this architecture.There are 364 sequences with the following architecture: Pkinase, POLO_box x 2
Q55K81_CRYNB [Cryptococcus neoformans var. neoformans serotype D (strain B-3501A) (Filobasidiella neoformans)] Putative uncharacterized protein (827 residues) Show all sequences with this architecture.There are 358 sequences with the following architecture: Pkinase, EF-hand_7 x 2
A0BDK2_PARTE [Paramecium tetraurelia] Chromosome undetermined scaffold_100, whole genome shotgun sequence (463 residues) Show all sequences with this architecture.There are 336 sequences with the following architecture: cNMP_binding x 2, Pkinase
A0BE58_PARTE [Paramecium tetraurelia] Chromosome undetermined scaffold_101, whole genome shotgun sequence (823 residues) Show all sequences with this architecture.There are 336 sequences with the following architecture: Pkinase, Ribonuc_2-5A
IRLB_DICDI [Dictyostelium discoideum (Slime mold)] Probable serine/threonine-protein kinase irlB EC=2.7.11.1 (1448 residues) Show all sequences with this architecture.There are 306 sequences with the following architecture: Stress-antifung x 2, Pkinase
Q8W5E9_ORYSJ [Oryza sativa subsp. japonica (Rice)] Putative receptor-like protein kinase (816 residues) Show all sequences with this architecture.There are 299 sequences with the following architecture: Activin_recp, Pkinase
AVR2A_CHICK [Gallus gallus (Chicken)] Activin receptor type-2A EC=2.7.11.30 (513 residues) Show all sequences with this architecture.There are 295 sequences with the following architecture: Pkinase, DUF3543
ATG1_EMENI [Emericella nidulans (strain FGSC A4 / ATCC 38163 / CBS 112.46 / NRRL 194 / M139) (Aspergillus nidulans)] Serine/threonine-protein kinase atg1 EC=2.7.11.1 (935 residues) Show all sequences with this architecture.There are 244 sequences with the following architecture: Pkinase, AAA_16
Q3JW39_BURP1 [Burkholderia pseudomallei (strain 1710b)] Protein kinase domain protein (1359 residues) Show all sequences with this architecture.There are 240 sequences with the following architecture: PP2C_2, Pkinase
Q21IE0_SACD2 [Saccharophagus degradans (strain 2-40 / ATCC 43961 / DSM 17024)] Serine/threonine protein kinase (574 residues) Show all sequences with this architecture.There are 228 sequences with the following architecture: Pkinase, CK1gamma_C
Q9VEX2_DROME [Drosophila melanogaster (Fruit fly)] Gilgamesh, isoform A EC=2.7.11.- (422 residues) Show all sequences with this architecture.There are 217 sequences with the following architecture: Mad3_BUB1_I, Pkinase
Q8K2E5_MOUSE [Mus musculus (Mouse)] Budding uninhibited by benzimidazoles 1 homolog, beta (S. cerevisiae) (1052 residues) Show all sequences with this architecture.There are 213 sequences with the following architecture: DUF1908, Pkinase, PDZ
MAST1_HUMAN [Homo sapiens (Human)] Microtubule-associated serine/threonine-protein kinase 1 EC=2.7.11.1 (1570 residues) Show all sequences with this architecture.There are 204 sequences with the following architecture: C1_1 x 2, Pkinase, Pkinase_C
Q1MX43_MOUSE [Mus musculus (Mouse)] Protein kinase C, delta IV (559 residues) Show all sequences with this architecture.There are 202 sequences with the following architecture: PH, Pkinase, Pkinase_C
AKT2_HUMAN [Homo sapiens (Human)] RAC-beta serine/threonine-protein kinase EC=2.7.11.1 (481 residues) Show all sequences with this architecture.There are 200 sequences with the following architecture: PB1, Pkinase
Q2PFR6_MACFA [Macaca fascicularis (Crab-eating macaque) (Cynomolgus monkey)] Putative uncharacterized protein (448 residues) Show all sequences with this architecture.There are 179 sequences with the following architecture: Pkinase, FGE-sulfatase
Q3KMM4_CHLTA [Chlamydia trachomatis serovar A (strain HAR-13 / ATCC VR-571B)] Serine/threonine-protein kinase PKN1 EC=2.7.1.- (614 residues) Show all sequences with this architecture.There are 178 sequences with the following architecture: GUB_WAK_bind, Pkinase
Q8RVT2_ORYSA [Oryza sativa (Rice)] Protein kinase (718 residues) Show all sequences with this architecture.There are 177 sequences with the following architecture: DUF4071, Pkinase
Q7KSB7_DROME [Drosophila melanogaster (Fruit fly)] Protein kinase at 92B, isoform B EC=2.7.11.- (1196 residues) Show all sequences with this architecture.There are 175 sequences with the following architecture: Pkinase, PKK x 2
STK10_HUMAN [Homo sapiens (Human)] Serine/threonine-protein kinase 10 EC=2.7.11.1 (968 residues) Show all sequences with this architecture.There are 168 sequences with the following architecture: C1_1 x 2, C2, Pkinase, Pkinase_C
O01715_HYDVU [Hydra vulgaris (Hydra) (Hydra attenuata)] Protein kinase C EC=2.7.11.13 (670 residues) Show all sequences with this architecture.There are 160 sequences with the following architecture: Pkinase, PH_3
Q76BX2_ASTPE [Asterina pectinifera (Starfish) (Patiria pectinifera)] Phosphoinositide dependent kinase-1 (571 residues) Show all sequences with this architecture.There are 155 sequences with the following architecture: HR1 x 3, Pkinase, Pkinase_C
PKN3_MOUSE [Mus musculus (Mouse)] Serine/threonine-protein kinase N3 EC=2.7.11.13 (878 residues) Show all sequences with this architecture.There are 154 sequences with the following architecture: B_lectin, Pkinase
Q0JEU6_ORYSJ [Oryza sativa subsp. japonica (Rice)] Os04g0202500 protein (807 residues) Show all sequences with this architecture.There are 152 sequences with the following architecture: Death, Pkinase
IRAK2_MOUSE [Mus musculus (Mouse)] Interleukin-1 receptor-associated kinase-like 2 (622 residues) Show all sequences with this architecture.There are 150 sequences with the following architecture: LRRNT_2, LRR_8, Pkinase
RLK90_ARATH [Arabidopsis thaliana (Mouse-ear cress)] Probable inactive receptor kinase RLK902 (647 residues) Show all sequences with this architecture.There are 147 sequences with the following architecture: RWD, Pkinase x 3, tRNA-synt_His, HGTP_anticodon2
Q5BB34_EMENI [Emericella nidulans (strain FGSC A4 / ATCC 38163 / CBS 112.46 / NRRL 194 / M139) (Aspergillus nidulans)] eIF2 alpha kinase (Eurofung) (1552 residues) Show all sequences with this architecture.There are 145 sequences with the following architecture: C1_1 x 2, PH, Pkinase
KPCD3_MOUSE [Mus musculus (Mouse)] Serine/threonine-protein kinase D3 EC=2.7.11.13 (889 residues) Show all sequences with this architecture.There are 144 sequences with the following architecture: B_lectin, S_locus_glycop, Pkinase
Q10B65_ORYSJ [Oryza sativa subsp. japonica (Rice)] Putative uncharacterized protein OSJNBa0010E04.17 (797 residues) Show all sequences with this architecture.There are 144 sequences with the following architecture: DCX x 2, Pkinase
DCLK_CAEEL [Caenorhabditis elegans] Serine/threonine-protein kinase zyg-8 EC=2.7.11.1 (802 residues) Show all sequences with this architecture.There are 144 sequences with the following architecture: Pkinase, LANC_like
Q5UW30_STRTR [Streptococcus thermophilus] OsrE (779 residues) Show all sequences with this architecture.There are 141 sequences with the following architecture: Pkinase, Mst1_SARAH
STK3_HUMAN [Homo sapiens (Human)] Serine/threonine-protein kinase 3 EC=2.7.11.1 (491 residues) Show all sequences with this architecture.There are 140 sequences with the following architecture: Pkinase, EF-hand_7 x 2, EF-hand_8
A0C9G2_PARTE [Paramecium tetraurelia] Chromosome undetermined scaffold_16, whole genome shotgun sequence (510 residues) Show all sequences with this architecture.There are 137 sequences with the following architecture: Pkinase, POLO_box
PLK4_HUMAN [Homo sapiens (Human)] Serine/threonine-protein kinase PLK4 EC=2.7.11.21 (970 residues) Show all sequences with this architecture.Currently showing architectures 1 to 50, out of a total of 4512.
Pfam Clan
This family is a member of clan PKinase (CL0016), which has the following description:
This superfamily includes the Serine/Threonine- and Tyrosine- protein kinases as well as related kinases that act on non-protein substrates.
The clan contains the following 19 members:
ABC1 APH APH_6_hur Choline_kinase DUF1679 DUF2252 EcKinase Fructosamin_kin Kdo Kinase-like PIP49_C Pkinase Pkinase_Tyr Pox_ser-thr_kin RIO1 Seadorna_VP7 UL97 WaaY YrbL-PhoP_regAlignments
We store a range of different sequence alignments for families. As well as the seed alignment from which the family is built, we provide the full alignment, generated by searching the sequence database using the family HMM. We also generate alignments using four representative proteomes (RP) sets, the NCBI sequence database, and our metagenomics sequence database. More...
View options
We make a range of alignments for each Pfam-A family. You can see a description of each above. You can view these alignments in various ways but please note that some types of alignment are never generated while others may not be available for all families, most commonly because the alignments are too large to handle.
| Seed (54) |
Full (114309) |
Representative proteomes | NCBI (134432) |
Meta (5368) |
||||
|---|---|---|---|---|---|---|---|---|
| RP15 (23780) |
RP35 (38306) |
RP55 (53856) |
RP75 (65911) |
|||||
| Jalview | ||||||||
| HTML | ||||||||
| PP/heatmap | 1 | |||||||
| Pfam viewer | ||||||||
1Cannot generate PP/Heatmap alignments for seeds; no PP data available
Key:
 available,
available,
 not generated,
— not available.
not generated,
— not available.
Format an alignment
Download options
We make all of our alignments available in Stockholm format. You can download them here as raw, plain text files or as gzip-compressed files.
| Seed (54) |
Full (114309) |
Representative proteomes | NCBI (134432) |
Meta (5368) |
||||
|---|---|---|---|---|---|---|---|---|
| RP15 (23780) |
RP35 (38306) |
RP55 (53856) |
RP75 (65911) |
|||||
| Raw Stockholm | ||||||||
| Gzipped | ||||||||
You can also download a FASTA format file containing the full-length sequences for all sequences in the full alignment.
External links
MyHits provides a collection of tools to handle multiple sequence alignments. For example, one can refine a seed alignment (sequence addition or removal, re-alignment or manual edition) and then search databases for remote homologs using HMMER3.
HMM logo
HMM logos is one way of visualising profile HMMs. Logos provide a quick overview of the properties of an HMM in a graphical form. You can see a more detailed description of HMM logos and find out how you can interpret them here. More...

Trees
This page displays the phylogenetic tree for this family's seed alignment. We use FastTree to calculate neighbour join trees with a local bootstrap based on 100 resamples (shown next to the tree nodes). FastTree calculates approximately-maximum-likelihood phylogenetic trees from our seed alignment.
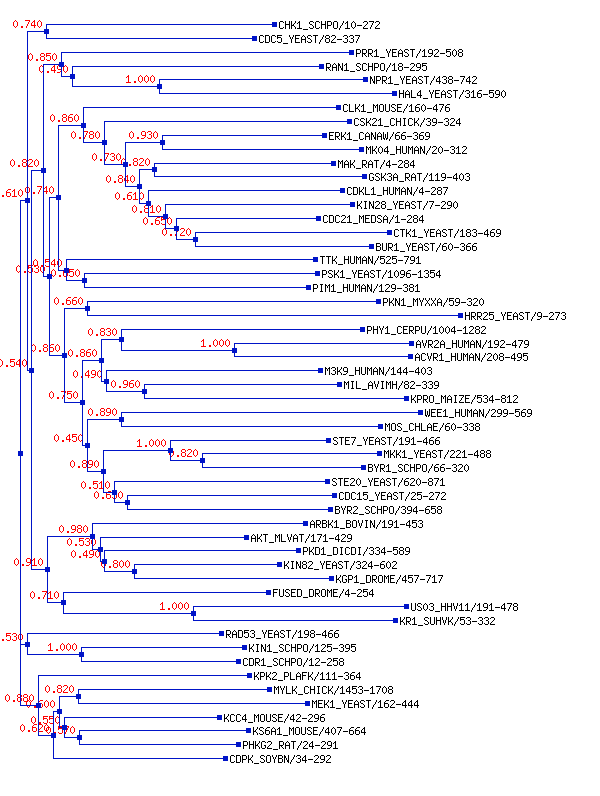
Note: You can also download the data file for the tree.
Curation and family details
This section shows the detailed information about the Pfam family. You can see the definitions of many of the terms in this section in the glossary and a fuller explanation of the scoring system that we use in the scores section of the help pages.
Curation

| Seed source: | Unknown |
| Previous IDs: | pkinase; |
| Type: | Domain |
| Author: | Sonnhammer ELL |
| Number in seed: | 54 |
| Number in full: | 114309 |
| Average length of the domain: | 237.60 aa |
| Average identity of full alignment: | 20 % |
| Average coverage of the sequence by the domain: | 40.43 % |
HMM information

| HMM build commands: |
build method: hmmbuild -o /dev/null HMM SEED
search method: hmmsearch -Z 23193494 -E 1000 --cpu 4 HMM pfamseq
|
||||||||||||
| Model details: |
|
||||||||||||
| Model length: | 260 | ||||||||||||
| Family (HMM) version: | 20 | ||||||||||||
| Download: | download the raw HMM for this family |
Species distribution
Sunburst controls
HideLineage
Move your mouse over the main tree to show the lineage of a particular node.
You can move this pane by dragging it.
Weight segments by...
Change the size of the sunburst
Colour assignments
 Archea
Archea
|
 Eukaryota
Eukaryota
|
 Bacteria
Bacteria
|
 Other sequences
Other sequences
|
 Viruses
Viruses
|
 Unclassified
Unclassified
|
 Viroids
Viroids
|
 Unclassified sequence
Unclassified sequence
|
Selections
Align selected sequences to HMM
Generate a FASTA-format file
Clear selection
This visualisation provides a simple graphical representation of the distribution of this family across species. You can find the original interactive tree in the adjacent tab. More...
Tree controls
HideThe tree shows the occurrence of this domain across different species. More...
Loading...
Please note: for large trees this can take some time. While the tree is loading, you can safely switch away from this tab but if you browse away from the family page entirely, the tree will not be loaded.
Interactions
There are 21 interactions for this family. More...
CDI UBA S1 Ank TGF_beta_GS K-cyclin_vir_C Cyclin_N RGS FKBP_C Pkinase_C Cyclin Herp-Cyclin Cyclin_C CK_II_beta Ribonuc_2-5A I-set cNMP_binding Pkinase PBD CDKN3 CKSStructures
For those sequences which have a structure in the Protein DataBank, we use the mapping between UniProt, PDB and Pfam coordinate systems from the PDBe group, to allow us to map Pfam domains onto UniProt sequences and three-dimensional protein structures. The table below shows the structures on which the Pkinase domain has been found. There are 2311 instances of this domain found in the PDB. Note that there may be multiple copies of the domain in a single PDB structure, since many structures contain multiple copies of the same protein seqence.
